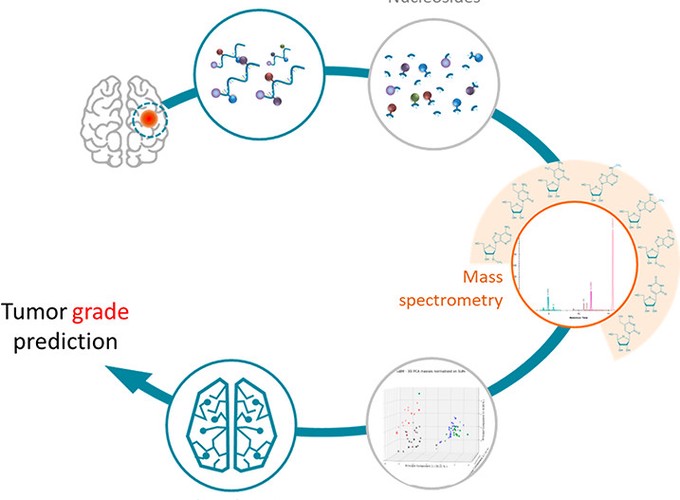
In a consortium allying a Mass Spectrometry platform, a cancer biology lab, several clinicians, and a bioinformatic team, we investigated how epigenetic marks on the RNA (a.k.a., the epitranscriptome) can help distinguishing the different grades of glioma tumors from tissue samples. We came up with a pipeline that proceeds surgical samples, measures the level of epitranscriptomic marks, and predicts the grade of this cancer. Our method delivers grade prediction with unmet sensitivity and specificity.
This research effort combines several disciplines: chemistry, cancer biology, and computer science. It was supported by the Occitanie region which funded the Mass Spectrometry platform for clinical purposes, known as the SMART project.
For more details, check out our publication in Analytical Chemistry entitled:
Multivariate Analysis of RNA Chemistry Marks Uncovers Epitranscriptomics-Based Biomarker Signature for Adult Diffuse Glioma Diagnostics
Authors: S. Relier, A. Amalric, A. Attina, I.B. Koumare, V. Rigau, F. Burel Vandenbos, D. Fontaine, M. Baroncini, J.P. Hugnot, H. Duffau, L. Bauchet, C. Hirtz*, E. Rivals*, and A. David* – *: co-corresponding authors.
Anal. Chem. 2022, doi: https://doi.org/10.1021/acs.analchem.2c01526