Overview
The way a cell controls how genes are activated to produce RNA and proteins relies on a complex, multi-step process called gene expression. It involves several levels of controls the two major ones being the transcription from DNA into RNA and the translation from RNA into protein. Gene expression is a key to understand cell differentiation and plays important roles during development or in diseases like cancer. Our project aims at investigating the mechanism and dynamics of transcription regulation and of translation. To address such complex issues, we will combine the modeling capacity of biophysics to bioinformatics data coming from genome wide analyses. Our goal is to design bioinformatic algorithms and models from statistical biophysics to understand how molecular players interact with polymers to control transcription or translation. Our main task is to launch a new type of multidisciplinary collaboration in synergy between biology, biophysics and bioinformatics, and to establish a dynamic and synergistic community on this topic in Montpellier.
Results
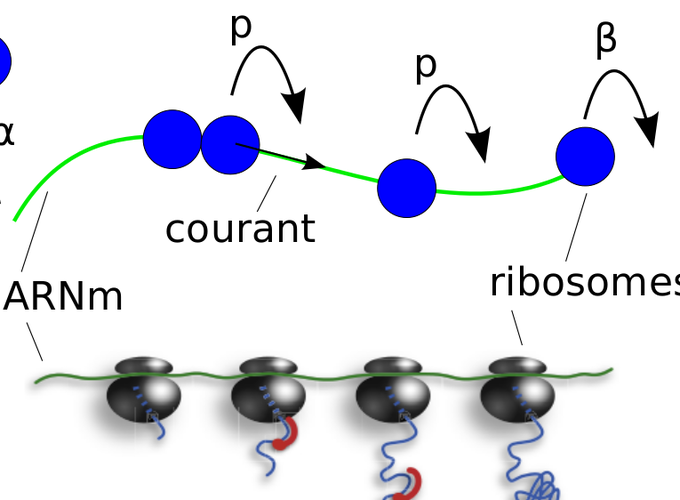
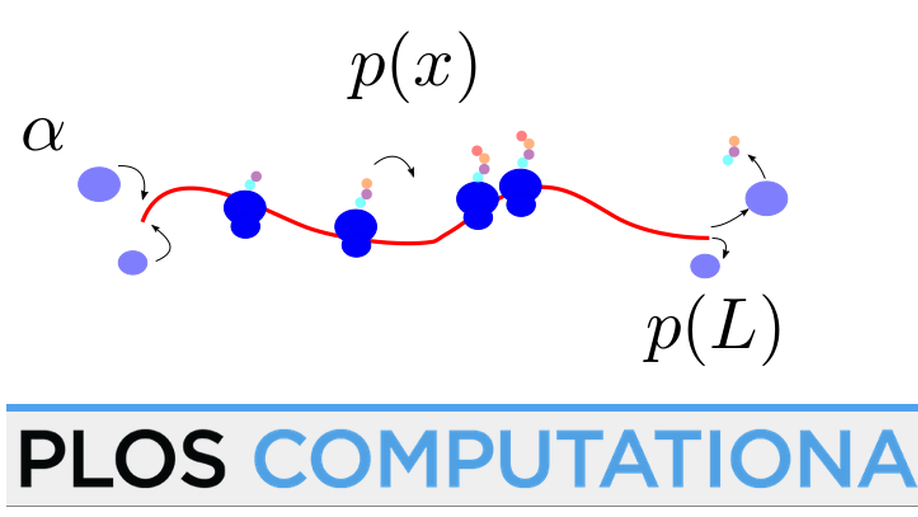
- Design of biophysical models to model the movement of ribosome during translation
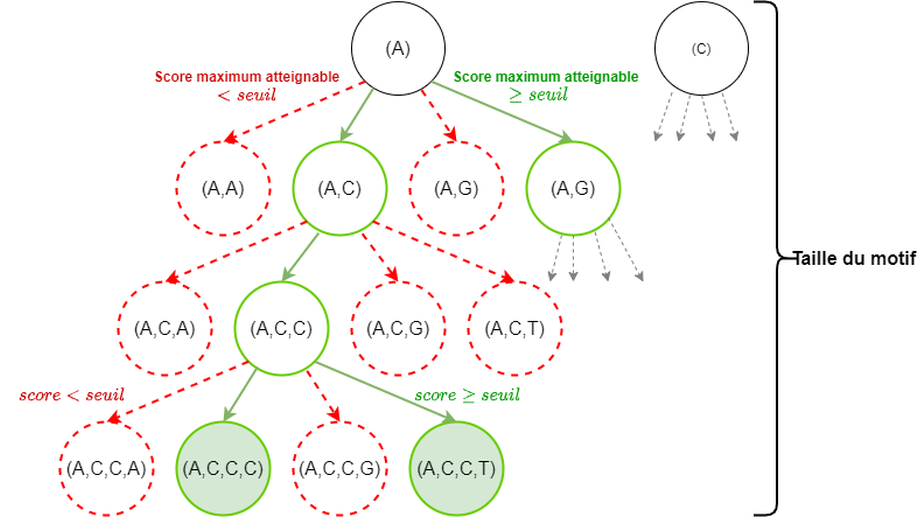
- Algorithms to process large datasets of DNA sequencing reads
- Study of a major player of RNA epigenetics